- Load the R packages we will use.
Download \(CO_2\) emissions per capita from Our World in Data into the directory for this post.
Assign the location of the file to
file.csv. The data should be in the same directory as this file.
- Read the data into R and assign it to
emissions
file_csv <- here("_posts",
"2021-02-16-reading-and-writing-data",
"co-emissions-per-capita.csv")
emissions <- read_csv(file_csv)
- Show the first 10 rows (observations of)
emissions
emissions
# A tibble: 22,383 x 4
Entity Code Year `Per capita CO2 emissions`
<chr> <chr> <dbl> <dbl>
1 Afghanistan AFG 1949 0.00191
2 Afghanistan AFG 1950 0.0109
3 Afghanistan AFG 1951 0.0117
4 Afghanistan AFG 1952 0.0115
5 Afghanistan AFG 1953 0.0132
6 Afghanistan AFG 1954 0.0130
7 Afghanistan AFG 1955 0.0186
8 Afghanistan AFG 1956 0.0218
9 Afghanistan AFG 1957 0.0343
10 Afghanistan AFG 1958 0.0380
# ... with 22,373 more rowsStart with
emissionsdata THENuse
clean namesfrom the janitor package to make the names easier to work withassign the output to
tidy_emissionsshow the first 10 rows of
tidy_emissions
tidy_emissions <- emissions %>%
clean_names()
tidy_emissions
# A tibble: 22,383 x 4
entity code year per_capita_co2_emissions
<chr> <chr> <dbl> <dbl>
1 Afghanistan AFG 1949 0.00191
2 Afghanistan AFG 1950 0.0109
3 Afghanistan AFG 1951 0.0117
4 Afghanistan AFG 1952 0.0115
5 Afghanistan AFG 1953 0.0132
6 Afghanistan AFG 1954 0.0130
7 Afghanistan AFG 1955 0.0186
8 Afghanistan AFG 1956 0.0218
9 Afghanistan AFG 1957 0.0343
10 Afghanistan AFG 1958 0.0380
# ... with 22,373 more rowsStart with the
tidy_emissionsTHENuse
filterto extract rows withyear==1985THENuse
skimto calculate the descriptive statistics
| Name | Piped data |
| Number of rows | 209 |
| Number of columns | 4 |
| _______________________ | |
| Column type frequency: | |
| character | 2 |
| numeric | 2 |
| ________________________ | |
| Group variables | None |
Variable type: character
| skim_variable | n_missing | complete_rate | min | max | empty | n_unique | whitespace |
|---|---|---|---|---|---|---|---|
| entity | 0 | 1.00 | 4 | 32 | 0 | 209 | 0 |
| code | 12 | 0.94 | 3 | 8 | 0 | 197 | 0 |
Variable type: numeric
| skim_variable | n_missing | complete_rate | mean | sd | p0 | p25 | p50 | p75 | p100 | hist |
|---|---|---|---|---|---|---|---|---|---|---|
| year | 0 | 1 | 1985.00 | 0.00 | 1985.00 | 1985.00 | 1985.00 | 1985.00 | 1985.00 | ▁▁▇▁▁ |
| per_capita_co2_emissions | 0 | 1 | 5.53 | 8.94 | 0.04 | 0.51 | 2.65 | 7.62 | 83.83 | ▇▁▁▁▁ |
12 observations have a missing code. How are these observations different?
- start with
tidy_emissionsthen extract rows withyear == 1985and are missing a code
- start with
# A tibble: 12 x 4
entity code year per_capita_co2_emissions
<chr> <chr> <dbl> <dbl>
1 Africa <NA> 1985 1.23
2 Asia <NA> 1985 1.81
3 Asia (excl. China & India) <NA> 1985 2.76
4 EU-27 <NA> 1985 9.19
5 EU-28 <NA> 1985 9.28
6 Europe <NA> 1985 10.9
7 Europe (excl. EU-27) <NA> 1985 13.3
8 Europe (excl. EU-28) <NA> 1985 14.1
9 North America <NA> 1985 13.2
10 North America (excl. USA) <NA> 1985 5.01
11 Oceania <NA> 1985 10.8
12 South America <NA> 1985 1.87Entities that are not countries do not have country codes.
Start with
tidy_emissionsTHENuse
filterto extract rows withyear==1985and without missing codes THENuse
selectto drop theyearvariable THENuse
renameto change the variableentitytocountryassign the output to
emissions_1985
Which 15 countries have the highest
per_capita_co2_emissions?start with emissions_1985 THEN
use
slice_maxto extract the 15 rows with the highest valuesassign the output to
max_15_emitters
max_15_emitters <- emissions_1985 %>%
slice_max(per_capita_co2_emissions, n=15)
Which 15 countries have the lowest
per_capita_co2_emissions?start with emissions_1985 THEN
use
slice_minto extract the 15 rows with the lowest valuesassign the output to
min_15_emitters
min_15_emitters <- emissions_1985 %>%
slice_min(per_capita_co2_emissions, n=15)
Use
bind_rowsto bind together themax_15_emittersandmin_15_emitters- assign the output to
max_min_15
- assign the output to
max_min_15 <- bind_rows(max_15_emitters, min_15_emitters)
- Export
max_min_15to 3 file formats
max_min_15 %>% write_csv("max_min_15.csv") # comma-separated values
max_min_15 %>% write_tsv("max_min_15.tsv") # tab separated
max_min_15 %>% write_delim("max_min_15.psv", delim = "|") #pipe-separated
- Read the 3 file formats into R
max_min_15_csv <- read_csv("max_min_15.csv") # comma-separated values
max_min_15_tsv <- read_tsv("max_min_15.tsv") # tab separated
max_min_15_psv <- read_delim("max_min_15.psv", delim = "|") #pipe-separated
- Use
setdiffto check for any differences amongmax_min_15_csv,max_min_15_tsvandmax_min_15_psv
setdiff(max_min_15_csv, max_min_15_tsv, max_min_15_psv)
# A tibble: 0 x 3
# ... with 3 variables: country <chr>, code <chr>,
# per_capita_co2_emissions <dbl>Are there any differences?
Reorder
countryinmax_min_15for plotting and assign to max_min_15_plot_datastart with emissions_1985 THEN
use
mutateto reordercountryaccording toper_capital_co2_emissions
max_min_15_plot_data <- max_min_15 %>%
mutate(country=reorder(country, per_capita_co2_emissions))
- Plot
max_min_15_plot_data
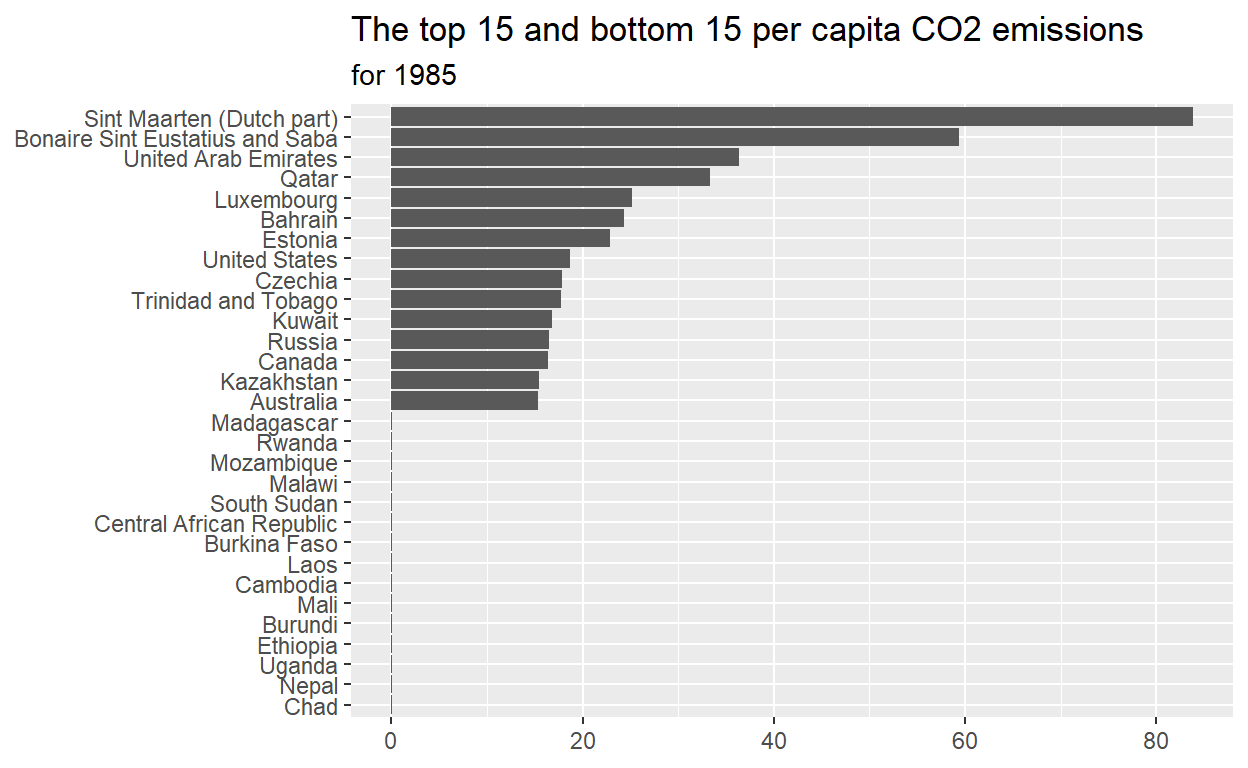
ggplot(data = max_min_15_plot_data ,
mapping=aes(x=per_capita_co2_emissions, y=country))+
geom_col()+
labs(title="The top 15 and bottom 15 per capita CO2 emissions",
subtitle = "for 1985",
x=NULL,
y=NULL)
- Save the plot directory with this post
ggsave(filename = "preview.png",
path=here("_posts", "2021-02-16-reading-and-writing-data"))
- Add preview.png to yaml chunk at the top of this file
preview:preview.png